Jan K. Rainey
Professor

Email: jan.rainey@dal.ca
Phone: 902-494-4632 (office)
Phone: 902-494-4812 (lab)
Mailing Address:
Sir Charles Tupper Medical Building
PO Box 15000
Halifax, Nova Scotia, Canada B3H 4R2
Education
- PhD, University of Toronto
Academic Positions
- Department member since 2006
- Cross-appointed to Department of Chemistry
- Cross-appointed to School of Biomedical Engineering
- Undergraduate Coordinator and Undergraduate Advisor, 2022-present
- Member, Protein Assembly Research Team
- Member, BioActives CREATE Training Program
Research Topics
Structural Biology and Biophysical Characterization of Proteins
Research
Structural & biophysical studies of membrane proteins & fibrous proteins
Proteins in the cell membrane and in fibres function in highly organized supramolecular assemblies, groupings of many molecules. Both classes of protein are critical for life, but are notoriously difficult to study. Our research goal is to determine not only how these classes of proteins function, but to carry out studies of these proteins in highly physiologically relevant environments. The primary aim is to understand the contribution of each constituent atom to the biological activity of a protein, providing fundamental understanding of biochemical processes, new routes for drug design, and alternative approaches for disease diagnosis and treatment. Current systems under study include a G-protein coupled receptor and its peptidic ligands; recombinant spider silk (one of the toughest known materials!); the extracellular matrix protein collagen (the most abundant protein in animals); soft polymeric nanoparticles; and, fusion-associate small transmembrane (FAST) proteins (a recently discovered class of viral membrane fusion proteins.)
Protein preparation
Two major approaches are used in our laboratory to produce proteins. In the organic chemistry based solid-phase peptide synthesis, a polypeptide is built by adding each amino acid in sequence. This provides us the ability to alter or tailor any individual amino acid residue we wish, either to determine effects of substitutions or to allow highly specific stable-isotope for nuclear magnetic resonance (NMR) spectroscopy or fluorescent labelling for biophysical study. Alternatively, molecular biology based cloning and expression is used to produce larger proteins. Beyond production of proteins, this readily allows both uniform and selective stable-isotope labelling . In each case, purification may be performed by high-performance liquid chromatography (HPLC) or other chromatographic methods, as appropriate. Mass spectrometry is typically used to characterize our synthetic or expression products.
NMR spectroscopy & structure calculations
NMR spectroscopy is our primary experimental technique, allowing us to determine protein structure and dynamics at the level of the individual nucleus. We use both solution-state and solid-state NMR to understand protein function in a variety of situations ranging from polypeptides in aqueous buffer to large protein assemblies and proteins embedded in phospholipid bilayers. In the case of solid-state NMR, we are currently working towards new methods that will allow characterization of not only the structure of proteins in membranes or fibres but the structure of these proteins relative to their supramolecular environment. Structure calculations are a critical component of biomolecular NMR, and we spend a lot of time ensuring that our calculations are truly providing a structural ensemble accurately representing the experimental data.
Scanning probe microscopy & force spectroscopy
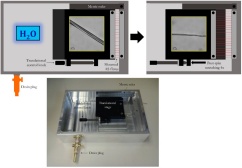
Scanning probe microscopy (SPM), also known as atomic force microscopy (AFM), is the other major technique employed by our laboratory. This type of microscopy uses a very sharp tip to image a surface, making use of the force of interaction between the tip and the surface to produce a topological and/or chemical image of the surface. This technique is capable of probing individual atoms, but for biological samples a more realistic resolution limit is on the order of a nanometer. SPM can also be used for study of the interaction forces between pairs of molecules, providing picoNewton level accuracy. A major focus in our research is the development of methods to allow exactly the same sample to be studied by solid-state NMR and SPM.
Circular dichroism & fluorescence spectroscopy
Circular dichroism (CD) spectroscopy allows rapid determination of protein secondary structure under various conditions. Fluorescence spectroscopy allows us to quickly assess the environment surrounding individual amino acids (i.e. are they exposed to solvent vs. buried in a protein core?) and the free- or bound-state of a protein. These methods provide an important counterpoint to our NMR and SPM experiments, since characterization is often much more rapidly achieved and provide different classes of information.
Funding
Operating: NSERC, CIHR, NSHRF. Infrastructure: CFI, NSERC, Dalhousie Medical Research Foundation.
Keywords:
Protein NMR spectroscopy, Solid-state NMR spectroscopy, scanning probe microscopy, circular dichroism (CD) spectroscopy, fluorescence spectroscopy, solid-phase peptide synthesis, cloning & expression, cell-free expression, HPLC, Langmuir-Blodgettry
Current Lab Members
| Suad Rashid | Postdoctoral Fellow |
|---|---|
| Yanitza Trosel | Postdoctoral fellow |
| Sara Evans | Grad Student (PhD Chemistry) |
| Hina Batool | Grad Student (PhD) |
| Oscar Alferez | Grad Student (PhD) |
| Isabela Drozdowski | Grad student (MSc; primary supervisor D. Langelaan)
|
| Andrew Chen | Summer & Honours Student (2025-2026) |
| Jessiny Dowell | Summer & Honours Student (2025-2026) |
| Charlotte Polo | Summer & Honours Student (2025-2026) |
| Woobeen Shin | Summer & Honours Student (2025-2026) |
| Ameeta Thandi | Summer & Honours Student (2025-2026) |
| Vivian Tia | Summer & Honours Student (2025-2026) |
| Noelle Aldous | BIOC 3620 student (2026) |
| Nikol Kot | BIOC 3620 student (2026) |
| Georgia McLenehan | BIOC 3620 student (2026) |
| Nora Meleschuk | BIOC 3620 student (2026) |
| Sofia Mereshuk | BIOC 3620 student (2026) |
| Theresa Martin | Volunteer researcher |
Publications
- Ghimire, A., Lee., S.Y., Chen, A.A.Y., Trosel, Y., Frampton, J.P., and Rainey, J.K. (2026) A bioengineered recombinant spider silk provides targeted G-protein-coupled receptor activation. Acta Biomater. In press. [PubMed] [Article]
- Ghimire, A., Chong, N.E.Y., Evans, S., Chen, A.A.Y., and Rainey, J.K., (2026) Recombinant pyriform-aciniform spidroin blends form enhanced silk fibers without discernible pre-assembly interactions. Int J Biol Macromol. 350: 150912. [PubMed] [Article]
- Kumar, S., Rainey, J.K., Stefanova, R., Zhang, J., and Brooks, M.S-L. (2026) Interactions and complex formation between pea proteins (Pisum sativum L.) and betanin from red beet (Beta vulgaris L.) extract – effect of foam fractionation. Food Hydrocoll. 172: 112139. [PubMed] [Article]
- Nazeer, N., Ghimire, A., Rainey, J.K., Lubell, W.D., Wagner, B.D., and Ahmed, M. (2026) Intrinsically fluorescent nano-scaled peptide aggregates upon arginine to citrulline swap. Biotechnol Bioeng. 123: 43-51 [PubMed] [Article]
- Mallett, T., Lamer, T., Aleksandrzak-Piekarczyk, T., McKay, R.T., Catenza, K., Sit, C., Rainey, J.K., Towle-Straub, K.M., Vederas, J.C., and van Belkum, M.J. (2025) Solution structure of the broad-spectrum bacteriocin garvicin Q. Int J Mol Sci. 26: 7846. [PubMed] [Article]
- Ghimire, A., Batool, H., Evans, S., Rashid, S., Rainey, D.L., Chen, A.A.Y., Liu, X-Q., and Rainey, J.K. (2025) Optimizing and evaluating hand-drawing and wet-spinning for recombinant spider silk fiber production. J Vis Exp. 221: e68714. [PubMed] [Article]
- Kumar, S., Rainey, J.K., Stefanova, R., Zhang, J., and Brooks, M.S-L. (2025) Interaction of pea protein isolate with betanin in red beet (Beta vulgaris L.) extract: Influence on structure, antioxidant properties and stability at pH 3. Food Chem. 492: 145419. [PubMed] [Article]
- Harvey, M., Ghimire, A., Cisek, R., Kreplak, L., Rainey, J.K., and Tokarz, D. (2025) Polarization-resolved second harmonic generation microscopy of silk fibers is sensitive to β-sheet orientation and molecular structure. ACS Appl Bio Mat. 8: 5229-5238 [PubMed] [Article]
- Sulekha, A., Evans, S., Badichi Akher, F., Ghimire, A., Reith, M.A., Liu, X-Q., Xu, L., and Rainey, J.K. (2025) An engineered recombinant spider silk protein providing disulfide-locked control over self-assembly and fiber formation. Adv Funct Mater. 35: 2420254. [Article]
- Nazeer, N., Kooner, N., Ghimire, A., Rainey, J.K., Lubell, W.D., Meneksedag Erol, D., and Ahmed, M. (2024) Secondary structure stabilization of macrocyclic antimicrobial peptides via cross-link swapping. J Med Chem. 67: 8693-8707. PMID: 38771638. [PubMed] [Article]
- Ghimire, A., Pham, T.T.T., Reith, M.A., and Rainey, J.K. (2024) Insight into the molecular basis of the soluble storage-state to fibrous-state structural transformation of spider pyriform silk. Can J Chem. 102: 802-812. [Article]
- Ghimire, A., Xu, L., Liu, X-Q., and Rainey, J.K. (2024) A recombinant chimeric spider pyriform-aciniform silk with highly tunable mechanical performance. Mater Today Bio. 26: 101073. PMID: 38711935. [PubMed] [Article]
- Reith, M.A. and Rainey, J.K. (2024) The critical importance of conditions: Reconciling GPCR functionality and biophysical findings. Structure. 32: 517-519. PMID: 38701749. [PubMed] [Article]
- Phạm, T.T.T., Murza, A., Marsault, É., Frampton, J.P., and Rainey, J.K. (2024) Localized apelin-17 analogue-bicelle interactions as a facilitator of membrane-catalyzed receptor recognition and binding. Biochim Biophys Acta Biomembr. 1866: 184289. [PubMed] [Article]
- Jahan, S., Doyle, C., Ghimire, A., Combita, D., Rainey, J.K., Wagner, B.D., and Ahmed, M. (2024) Elucidating the role of optical activity of polymers in protein-polymer interactions. Polymers. 16: 65. [Article]
- Kumar, P., Kermanshahi-Pour, A., Brar, S.K., Xu, C.C., He, Q.S., Evans, S., and Rainey, J.K. (2023) Enzymatic digestibility of lignocellulosic wood biomass: Effect of enzyme treatment in supercritical carbon dioxide and biomass pretreatment. Heliyon. 9: e21811. PMID: 38027598 [PubMed] [Article]
- Kumar, P., Kermanshahi-Pour, A., Brar, S.K., He, Q.S., and Rainey, J.K. (2023) Influence of elevated pressure and pressurized fluids on microenvironment and activity of enzymes. Biotechnol Adv. 68: 108219. PMID: 37488056. [PubMed] [Article]
- Simmons, J.R., Gasmi-Seabrook, G., and Rainey, J.K. (2023) Structural features, intrinsic disorder, and modularity of a pyriform spidroin 1 core repetitive domain. Biochem Cell Biol. 101: 271-283 [PubMed] [Article]
- Song, Q., Liu, X-Q., and Rainey, J.K. (2023) 1H, 15N and 13C backbone resonance assignments of the acidic domain of the human MDM2 protein. Biomol NMR Assign. 17: 9-16. [PubMed] [Article]
- Rajan, V., Prykhozhij, S.V., Pandey, A., Cohen, A.M., Rainey, J.K., and Berman, J.N. (2023) KIT D816V is dimerization-independent and activates downstream pathways frequently perturbed in mastocytosis. British Journal of Haematology. 202: 960-970. [PubMed] [Article]
- Visser, Z.B., Verma, S.K., Rainey, J.K., and Frampton, J.P. (2022) Loading and release of quercetin from contact-drawn polyvinyl alcohol fiber scaffolds. ACS Pharmacol Transl Sci. 5: 1305-1317. [PubMed] [Article]
- Baker, L.A., Xu, L., Badichi Akher, F., Robertson, M.K., Pugsley-DeBruyn, L., Ma, C.X., Liu, X-Q., Frampton, J.P., and Rainey, J.K. (2022) Nerve growth factor-binding engineered silk films promote neuronal attachment and neurite outgrowth. Adv Funct Mater. 3:2205178. [Article]
- Song, Q., Liu, X-Q., and Rainey, J.K. (2022) The MDMX acidic domain competes with the p53 transactivation domain for MDM2 N-terminal domain binding. Biochim Biophys Acta Mol Cell Res. 1869: 119319. [PubMed] [Article]
- Verma, S.K., Yaghoobi, H., Slaine, P., Baldwin, S.K., Rainey, J.K., Kreplak, L., and Frampton, J.P. (2022) Multi-pin contact drawing enables production of anisotropic collagen fiber substrates for alignment of fibroblasts and monocytes. Colloids Surf B. 215:112525. [Article]
- Song, Q., Liu, X-Q., and Rainey J.K. (2022) 1H, 15N and 13C backbone resonance assignments of the acidic domain of the human MDMX protein. Biomol NMR Assign. 16:171-178. [PubMed] [Article]
- Dagamajalu, S., Rex, D.A.B., Suchitha, G.P., Rai, A.B., Rainey, J.K., and Prasad, T.S.K. (2022) The network map of Elabela signaling pathway in physiological and pathological conditions. J Cell Commun Signal. 16: 145-154. [PubMed] [Article]
- Dagamajalu, S., Rex, D.A.B., Philem, P.D., Rainey, J.K., and Keshava Prasad, T.S. (2022) A network map of apelin-mediated signaling. J Cell Commun Signal. 16:137-143. [PubMed] [Article]
- Nazeer, N., Simmons, J. R., Rainey, J. K., Rodriguez-Lecompte, J. C., & Ahmed, M. (2021). Antibacterial activities of physiologically stable, self-assembled peptide nanoparticles. J Mater Chem B. 9:9041-9054. [PubMed] [Article]
- Phạm, T.T.T. and Rainey, J.K. (2021) On-cell nuclear magnetic resonance spectroscopy to probe cell surface interactions. Biochem Cell Biol. 99:683-692. [PubMed] [Article]
- Tam, N.W., Chung, D., Baldwin, S.J., Simmons, J.R., Xu, L., Rainey, J.K., Dellaire, G., and Frampton, J.P. (2020) Material properties of disulfide-crosslinked hyaluronic acid hydrogels influence prostate cancer cell growth and metabolism. J. Mater. Chem. B. 8: 9718-9733. [PubMed] [Article]
- Shin, K., Landsman, M., Pelletier, S., Alamri, B., Anini, Y. and Rainey, J.K., (2019) Proapelin is processed extracellularly in a cell line-dependent manner with clear modulation by proprotein convertases Amino Acids 51:395-405 [PubMed] [Article]
- Sarker, M., Speckert, M. and Rainey, J.K., (2019) Bicelle composition-dependent modulation of phospholipid dynamics by apelin peptides. Biochem. Cell Biol. 97:325-332 [PubMed] [Article]
- Pandey, A., LeBlanc, D.M., Parmar, H.B., Phạm, T.T.T., Sarker, M., Xu, L., Duncan, R., Liu, X-Q. and Rainey, J.K., (2019) Structure, amphipathy, and topology of the membrane-proximal helix 8 influence apelin receptor plasma membrane localization Biochim Biophys Acta - Biomembranes 1861:180036 [PubMed] [Article]
- Xu, L., Weatherbee-Martin, N., Liu, X-Q. and Rainey, J.K., (2019) Recombinant silk fiber properties correlate to prefibrillar self-assembly Small 15:1805294 [PubMed] [Article]
- Simmons, J.R., Xu, L., and Rainey J.K., (2019) Recominant pyriform silk fiber mechanics are modulated by wet-spinning conditions. ACS Biomater Sci Eng 5:4985-4993 [Article]
- Getz, L.J., Runte, C.S., Rainey, J.K. and Thomas, N.A., (2019) Tyrosine phosphorylation as a widespread regulatory mechanism in prokaryotes J Bacteriol 201:e00205-19 [PubMed] [Article]
- Simmons, J.R., Murza, A., Lumsden, M.D., Kenward, K., Marsault, É. and Rainey J.K., (2019) Simultaneous ligand and receptor tracking through NMR spectroscopy enabled by distinct 19F labels. Int J Mol Sci 20:3658 [PubMed] [Article]
- Pandey, A.; Leblanc, D. M.; Parmar, H. B.; Pham, T. T. T.; Sarker, M.; Xu, L.; Duncan, R.; Liu, X. and Rainey, J. K. , (2019) Structure, amphipathy, and topology of the membrane-proximal helix 8 influence apelin receptor plasma membrane localization. BBA - Biomembranes 1861(11):183036 [PubMed] [Article]
- Shin, K., Kenward, C. and Rainey, J.K., (2018) Apelinergic system structure and function Compr Physiol 8:407-450 [PubMed] [Article]
- Kenward, C., Shin, K. and Rainey, J.K., (2018) Mixed Fluorotryptophan Substitutions at the Same Residue Expand the Versatility of 19F Protein NMR Spectroscopy Chem Eur J 24:3391-3396 [PubMed] [Article]
- Morash, B., Sarker, M. and Rainey, J.K., (2018) Concentration-dependent changes to diffusion and chemical shift of internal standard molecules in aqueous and micellar solutions J Biomol NMR 71:79-89 [PubMed] [Article]
- Huang, S.K., Shin, K., Sarker, M. and Rainey, J.K., (2017) Apela exhibits isoform- and headgroup-dependent modulation of micelle binding, peptide conformation and dynamics. Biochim Biophys Acta - Biomembranes 1859:767-778 [PubMed] [Article]
- Zhong, X.Z., Zou, Y., Sun, X., Dong, G., Cao, Q., Pandey, A., Rainey, J.K., Zhu, X. and Dong, X-P., (2017) Inhibition of TRPML1 by lysosomal adenosine involved in severe combined immunodeficiency diseases. J. Biol. Chem. 292:3445-3455 [PubMed] [Article]
- Young, B.M., Nguyen, E., Chedrawe, M.A.J., Rainey, J.K. and Dupré, D.J., (2017) Differential contribution of transmembrane domains IV, V, VI, and VII to human angiotensin II type 1 receptor homomer formation J. Biol. Chem. 292:3341-3350 [PubMed] [Article]
- Shin, K., Chapman, N.A., Sarker, M., Kenward, C., Huang, S.K., Weatherbee-Martin, N., Pandey, A., Dupré, D.J. and Rainey, J.K., (2017) Bioactivity of the putative apelin proprotein expands the repertoire of apelin receptor ligands. Biochim Biophys Acta - General Subjects 1861:1901-1912 [PubMed] [Article]
- Patterson, R.E., Weatherbee-Martin, N. and Rainey, J.K., (2017) Pyrene-Apelin Conjugation Modulates Fluorophore- and Peptide-Micelle Interactions. J Phys Chem B 121:47684777 [PubMed] [Article]
- Langelaan, D.N., Pandey, A., Sarker, M. and Rainey, J.K., (2017) Preserved transmembrane segment topology, structure, and dynamics in disparate micellar environments J Phys Chem Lett 8:2381-2386 [PubMed] [Article]
- Shin, K., Sarker, M., Huang, S.K., and Rainey, J.K., (2017) Apelin conformational and binding equilibria upon micelle interaction primarily depend on membrane-mimetic headgroup Sci Rep 7:15433 [PubMed] [Article]
- Xu, L., Lefèvre, T., Orrell, K.E., Meng, Q., Auger, M., Liu, X-Q. and Rainey, J.K., (2017) Structural and mechanical roles for the C-terminal non-repetitive domain become apparent in recombinant spider aciniform silk. Biomacromolecules 18:3678-3686 [PubMed] [Article]
- Das. D., Shink, K., Rainey, J.K. and Fliegel, L., (2017) Transmembrane segment XI of the Na+/H+ antiporter of S. pombe is a critical part of the ion translocation pore. Sci Rep 7:12793 [PubMed] [Article]
- Tremblay, M-L., Xu, L., Sarker, M., Liu, X-Q. and Rainey, J.K., (2016) Characterizing aciniform silk repetitive domain backbone dynamics and hydrodynamic modularity. Int. J. Mol. Sci. 17:E1305 [PubMed] [Article]
- Pandey, A., Shin, K., Patterson, R.E., Liu, X-Q. and Rainey, J.K., (2016) Current strategies for protein production and purification enabling membrane protein structural biology. Biochem. Cell Biol. 94(6):507-527 [PubMed] [Article]
- Weatherbee-Martin, N., Xu, L., Hupe, A., Kreplak, L., Fudge, D.S., Liu, X-Q. and Rainey, J.K., (2016) Identification of wet-spinning and post-spin stretching methods amenable to recombinant spider aciniform silk Biomacromolecules 17:2737-2746 [PubMed] [Article]
- Sarker, M., Orrell, K.E., Xu, L., Tremblay, M-L., Bak, J.J., Liu, X-Q. and Rainey, J.K., (2016) Tracking transitions in spider wrapping silk conformation and dynamics by 19F nuclear magnetic resonance spectroscopy Biochemistry 55:30483059 [PubMed] [Article]
- Read, J., Clancy, E.K., Sarker, M., de Antueno, R., Langelaan, D.N., Parmar, H.B., Shin, K., Rainey, J.K. and Duncan, R., (2015) Reovirus FAST proteins drive pore formation and syncytiogenesis using a novel helix-loop-helix fusion-inducing lipid packing sensor PLoS Pathog 11(6):e1004962 [PubMed] [Article]
- Sarker, M., Fraser, R.E., Lumsden, M.D., Anderson, D.J. and Rainey, J.K., (2015) Characterization of variant soft nanoparticle structure and morphology in solution by NMR spectroscopy. J Phys Chem C 119:7461-7471 [Article]
- Key, T., Sarker, M., de Antueno, R., Rainey, J.K. and Duncan, R., (2015) The p10 FAST protein fusion peptide functions as a cystine noose to induce cholesterol-dependent liposome fusion without liposome tubulation. Biochim Biophys Acta - Biomembranes 1848:408-416 [PubMed] [Article]
- Eftaiha, A.F., Tremblay, M-L., Rainey, J.K. and Paige, M.F, (2015) The effect of perfluorooctadecanoic acid on a model phosphatidylcholine-peptide pulmonary lung surfactant mixture. J. Fluorine Chem. 177:55-61 [Article]
- Clattenburg, L., Wigerius, M., Qi, J., Rainey, J.K., Rourke, J.L., Muruganandan, S., Sinal, C.J. and Fawcett, J.P., (2015) NOS1AP functionally associates with YAP to regulate Hippo signaling Mol. Cell. Biol. 35:2265-2277 [PubMed] [Article]
- Tremblay, M-L., Xu, L., Lefèvre, T., Sarker, M., Orrell, K.E., Leclerc, J., Meng, Q., Pezolet, M., Auger, M., Liu, X-Q. and Rainey, J.K., (2015) Spider wrapping silk fiber architecture arising from its modular soluble protein precursor. Sci. Rep. 5:11502 [PubMed] [Article]
- Pandey, A., Sarker, M., Liu, X-Q. and Rainey, J.K., (2014) Small expression tags enhance bacterial expression of the first three transmembrane segments of the apelin receptor. Biochem Cell Biol 92:269-278 [PubMed] [Article]
- Chapman, N.A., Dupré, D.J. and Rainey, J.K., (2014) The apelin receptor: Physiology, pathology, cell signalling, and ligand modulation of a peptide-activated class A GPCR. Biochem Cell Biol 92:431-440 [PubMed] [Article]
- Ramu, T., Prasad, M., Connors, E., Mishra, A., Thomassin, J-L., Leblanc, J., Rainey, J.K. and Thomas, N., (2013) A novel C-terminal region within the multicargo type III secretion chaperone CesT contributes to effector secretion. J. Bacteriol. 195:740-756 [PubMed]
- Xu, L., Tremblay, M-L., Orrell, K.E., Leclerc, J., Meng, Q., Liu, X-Q. and Rainey, J.K., (2013) Nanoparticle self-assembly by a highly stable recombinant spider wrapping silk protein subunit. FEBS Lett 587:3273-3280 [PubMed] [Article]
- Langelaan, D.N., Reddy, T., Banks, A.W., Dellaire, G., Dupré, D. and Rainey, J.K., (2013) Structural features of the apelin receptor N-terminal tail and first transmembrane segment implicated in ligand binding and receptor trafficking. Biochimica et Biophysica Acta - Biomembranes 1828:1471-1483 [PubMed] [Article]
- Kehoe, S., Tremblay, M-L., Coughlan, A., Towler, M.R., Rainey, J.K., Abraham, R.J. and Boyd, D., (2013) Preliminary Investigation of the Dissolution Behavior, Cytocompatibility, Effects of Fibrinogen Conformation and Platelet Activation for Radiopaque Embolic Particles. J. Funct. Biomater. 4:89-113 [Article]
- Shin, K., Pandey, A., Liu, X-Q., Anini, Y. and Rainey, J.K., (2013) Preferential apelin-13 production by the proprotein convertase PCSK3 is implicated in obesity. FEBS Open Bio. 3:328-333 [PubMed] [Article]
- Xu, L., Rainey, J.K., Meng, Q. and Liu, X-Q. , (2012) Recombinant minimalist spider wrapping silk proteins capable of native-like fiber formation. PLOS ONE 7:e50227 [PubMed]
- Xu, L.*, Tremblay, M-L.*, Meng, Q., Liu, X-Q. and Rainey, J.K. (* contributed equally), (2012) 1H, 13C and 15N NMR assignments of the aciniform spidroin (AcSp1) repetitive domain of Argiope trifasciata wrapping silk. Biomol. NMR Assign. 6:147-151 [PubMed]
- Reddy, T. and Rainey, J.K., (2012) The multifaceted substrate capture scheme of a rhomboid protease. J. Phys. Chem. B 116:8942-8954 [PubMed]
- Langelaan, D.N.*, Ngweniform, P.*, Rainey, J.K. (*contributed equally), (2011) Biophysical characterization of G-protein coupled receptor-peptide ligand binding. Biochem Cell Biol 89:98-105 [PubMed]
- Reddy, T., Li, X., Fliegel, L., Sykes, B.D. and Rainey, J.K., (2010) Correlating structure, dynamics, and function in transmembrane segment VII of the Na+/H+ exchanger isoform 1. Biochim Biophys Acta Biomembranes 1798:94-104 [PubMed]
- Langelaan, D.N. and Rainey, J.K., (2010) Membrane catalysis of peptide-receptor binding. Biochem Cell Biol 88:203-210 [PubMed]
- Reddy, T. and Rainey, J.K., (2010) Interpretation of biomolecular NMR spin relaxation parameters. Biochem Cell Biol 88:131-142 [PubMed]
- Tremblay, M-L., Banks, A.W. and Rainey, J.K., (2010) The predictive accuracy of secondary chemical shifts is more affected by protein secondary structure than solvent environment. J Biomol NMR 46:257-270 [PubMed]
- Langelaan, D.N., Wieczorek, M., Blouin, C. and Rainey, J.K., (2010) Improved helix and kink characterization in membrane proteins allows evaluation of kink sequence predictors. J. Chem. Inf. Model. 50:2213-2220 [PubMed]
- Langelaan, D.N., Bebbington, E.M., Reddy, T. and Rainey, J.K., (2009) Structural insight into G-protein coupled receptor binding by apelin Biochemistry 48:537-548 [PubMed]
- Langelaan, D.N. and Rainey, J.K., (2009) Headgroup-dependent membrane catalysis of apelin-receptor interactions is likely. J Phys Chem B 113:10465-10471 [PubMed]
- Reddy, T., Ding, J., Li, X., Sykes, B.D., Rainey, J.K. and Fliegel, L., (2008) Structural and functional characterization of transmembrane segment IX of the NHE1 isoform of the Na+/H+ exchanger. J. Biol. Chem. 283:22018-22030 [PubMed]